Reconstructing Queen Genotypes by Pool Sequencing Colonies in Eusocial Insects: Statistical Methods and Their Application to Honeybee
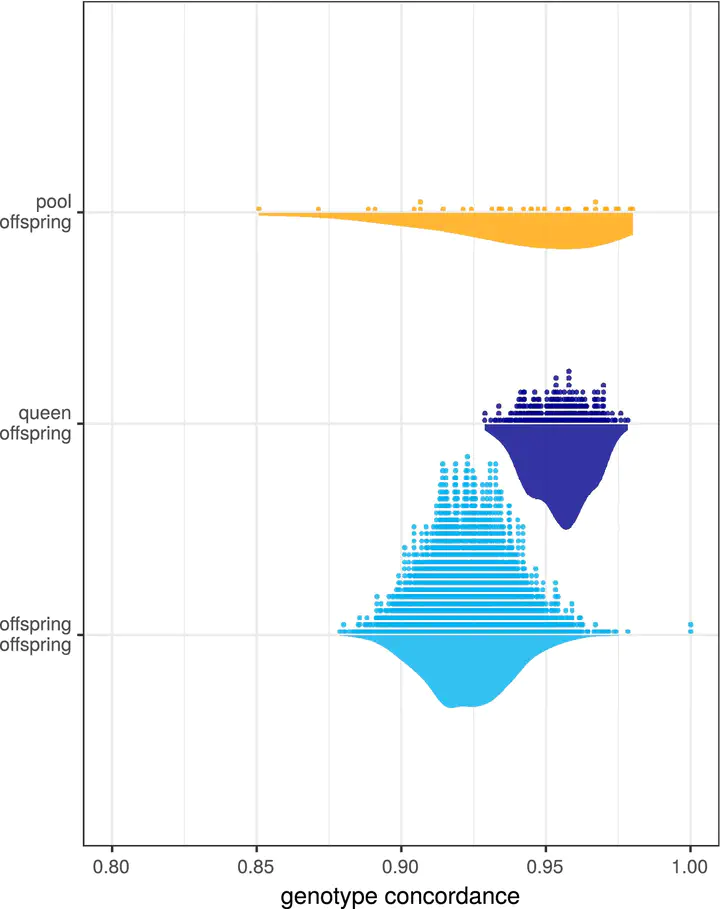 Concordance between queen genotypes reconstruction based on different data.
Concordance between queen genotypes reconstruction based on different data.Abstract
Eusocial insects are crucial to many ecosystems, and particularly the honeybee (Apis mellifera). One approach to facilitate their study in molecular genetics, is to consider whole-colony genotyping by combining DNA of multiple individuals in a single pool sequencing experiment. Cheap and fast, this technique comes with the drawback of producing data requiring dedicated methods to be fully exploited. Despite this limitation, pool sequencing data have been shown to be informative and cost-effective when working on random mating populations. Here, we present new statistical methods for exploiting pool sequencing of eusocial colonies in order to reconstruct the genotypes of the queen of such colony. This leverages the possibility to monitor genetic diversity, perform genomic-based studies or implement selective breeding. Using simulations and honeybee real data, we show that the new methods allow for a fast and accurate estimation of the queen’s genetic ancestry, with correlations of about 0.9 to that obtained from individual genotyping. Also, it allows for an accurate reconstruction of the queen genotypes, with about 2% genotyping error. We further validate these inferences using experimental data on colonies with both pool sequencing and individual genotyping of drones. In brief, in this study we present statistical models to accurately estimate the genetic ancestry and reconstruct the genotypes of the queen from pool sequencing data from workers of an eusocial colony. Such information allows to exploit pool sequencing for traditional population genetics analyses, association studies and for selective breeding. While validated in Apis mellifera, these methods are applicable to other eusocial hymenopterans.